Blocking Peptide Protocols for Immunohistochemistry (IHC) and Immunocytochemistry (ICC)
Improve antibody validation in IHC and ICC with a simple blocking peptide control.
A blocking peptide is the original antigen we use for immunization during antibody generation. As such, a blocking peptide is a great control to help validate the specificity of your antibody. Sometimes called “immunizing peptides” or “negative control antigens,” a blocking peptide works as a control by competing with, or blocking, the primary antibody.
You can easily use a blocking peptide control alongside your immunohistochemistry (IHC) and immunocytochemistry (ICC) experiments to help show that your antibody binds the intended target. It’s important to remember that a blocking peptide is just one of the many tools you should use for proper immunoassay controls.
An antibody blocked with a blocking peptide should produce no signal when added to your tissue sections. Any positive results in the blocking peptide control mean the antibody is binding to a protein besides the target.
Method
- Complete the preparation, fixation, and permeabilization (for intracellular epitopes) of your tissue section or cells according to the immunostaining protocol. Refer to our IHC, ICC, or IF protocols for further information.
- Reconstitute your lyophilized blocking peptide in 40 µl of sterile IHC/ICC phosphate-buffered saline (IHC/ICC-PBS) according to the instructions in the datasheet.
| IHC-PBS (pH 7.4) | ||
|---|---|---|
| Reagent | Concentration | Volume/Weight |
| Na2HPO4 | 0.2 M | 80 ml |
| KH2PO4 | 0.2 M | 16 ml |
| NaCl | 8 g | |
| Double distilled water | 860 ml | |
- Optimize your protocol in advance. Follow the product guidelines or previous experimental optimizations for a recommended dilution for your primary antibody.
- If your optimal antibody concentration is a 1:200 dilution, add 6 µl of antibody to an Eppendorf tube that contains 1.2 ml of Antibody Solution.
| Antibody Solution | |
|---|---|
| Reagent | % of final volume |
| IHC/ICC-PBS | 97.65 |
| Triton X-100* | 0.3 |
| Tween-20 | 0.05 |
| Normal serum** | 2 |
*If your primary antibody targets an extracellular protein, reduce the Triton X-100 to 0.05% in both the primary and secondary antibody solutions.
**Use a serum based on the species that your secondary antibody was raised in. For example, if your secondary antibody was raised in donkeys, use normal donkey serum (NDS). Likewise, if your secondary antibody was raised in goats, use normal goat serum (NGS).
- Divide the solution into two different identical Eppendorf tubes. The first one contains 600 µl of Antibody Solution with antibody. Label the tube “antibody alone”.
- To the second identical tube, add 40 µl of blocking peptide. Label the tube “+peptide”.
Note: we recommend that the concentration (mg/ml) of the blocking peptide be at least 10x the concentration of the antibody in the working dilution. - Rotate both tubes for 1 hour at room temperature.
- Add the contents of each tube to its respective tissue section well for parallel staining experiments.
- Incubate at room temperature for 1 hour with occasional gentle shaking of the multi-well plate.
- Incubate the multi-well plate overnight at 4°C.
- Proceed with the IHC, ICC, or IF protocol making sure you handle the unblocked and blocked tissues in the same way.
- Observe and compare the staining pattern obtained in the two test slides. The staining that disappears when using the blocking peptide is specific to the antibody. Any other staining that is visible represents non-specific binding.
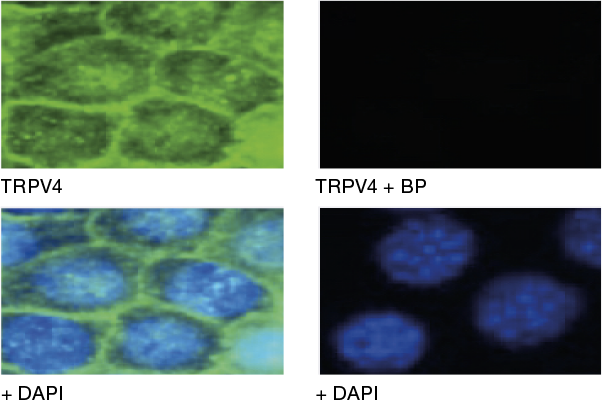
Example Data
Immunocytochemical staining using Anti-TRPV4 Antibody (ACC-034) within the presence or absence of blocking peptide. Immunocytochemical staining of mouse mCCDcl1 kidney cells using the Anti-TRPV4 Antibody (ACC-034) (panel TRPV4; green). The TRPV4 staining was completely abolished when the antibody was preincubated with the TRPV4 Blocking Peptide (BLP-CC034) (panel TRPV4 +BP). DAPI staining confirms the location of the nuclei.
Adapted from Li, Y. et al. (2016) PLoS ONE 11, e0155006 with permission from PLoS.